Improved graphical overview of data and results pertaining to the recently extended regression analysis and ANOVA
Prepare data
Dependence of on certain variables
Dependence on gene, age, and institution
Implementation of the same plot both with the lattice and the ggplot2 package.
P <- list()
# lattice implementation
P$s.age.inst$lattice <-
xyplot(S ~ Age | Gene, data = Y.long,
subset = Gene %in% gene.ids,
groups = Institution,
panel = function(x, y, ...) {
panel.xyplot(x, y, pch = 21, cex = 0.3, ...)
panel.smoother(x, y, col = "black", lwd = 2, ...)
},
auto.key = list(title = "institution", columns = 3),
par.settings = list(add.text = list(cex = 0.8)),
ylab = "read count ratio, S",
xlab = "age",
aspect = "fill", layout = c(6, 5))
# ggplot2 implementation
g <- ggplot(data = Y.long, aes(x = Age, y = S))
g <- g + geom_point(pch = "o", aes(color = Institution))
g <- g + geom_smooth(method = "loess", color = "black")
g <- g + facet_wrap(~ Gene)
P$s.age.inst$ggplot2 <- g
plot(P$s.age.inst$lattice)
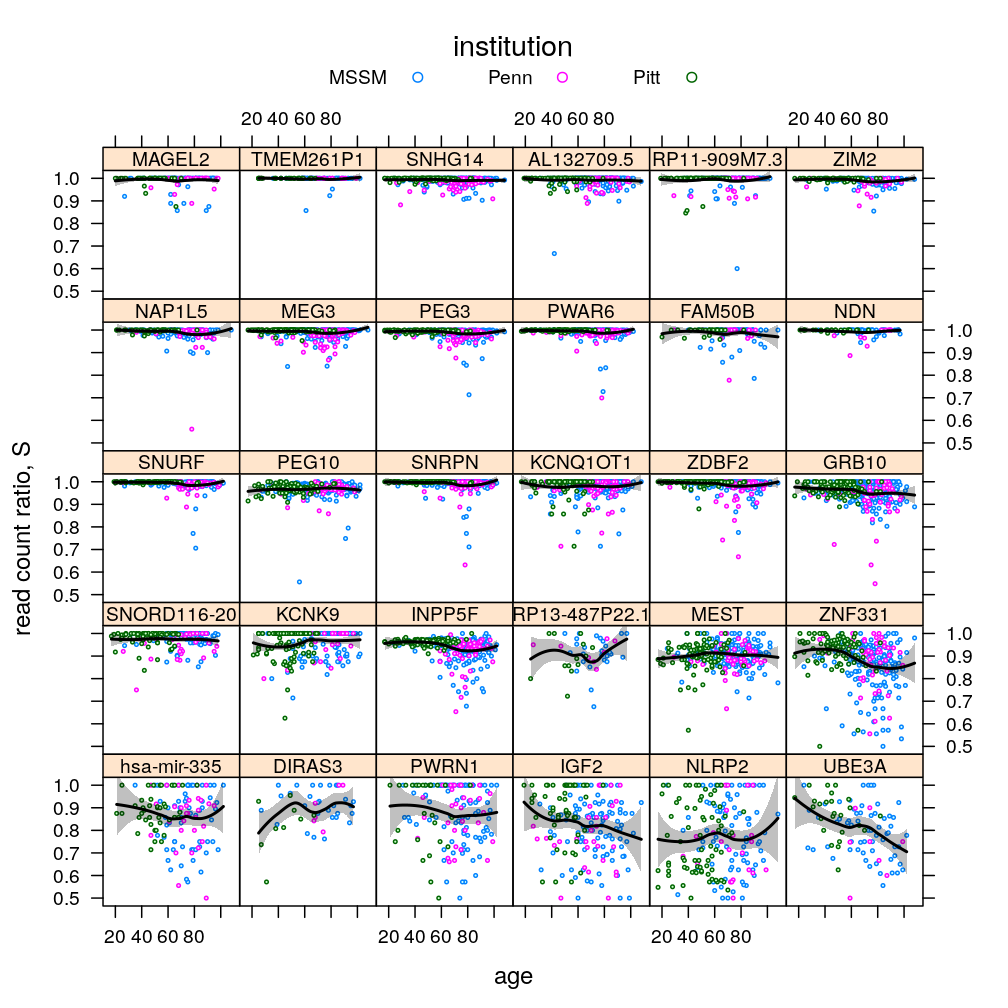
plot(P$s.age.inst$ggplot2)
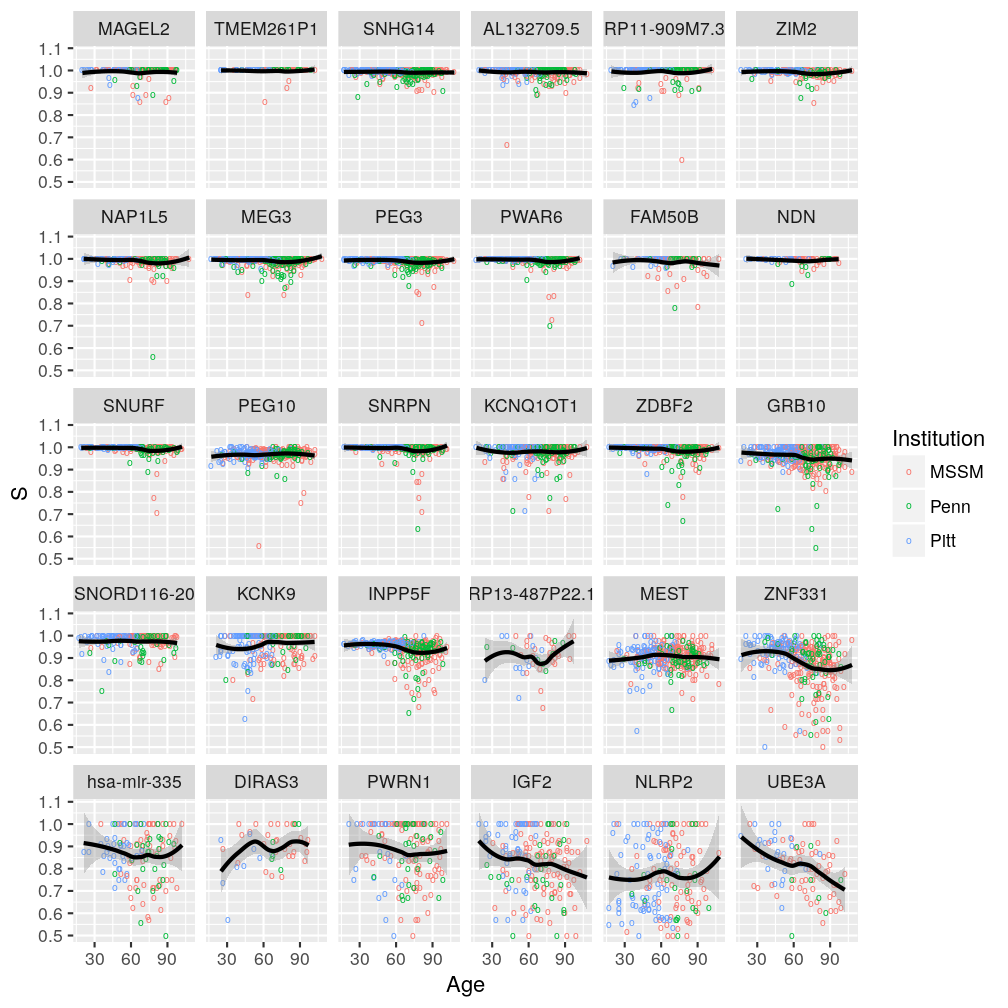
Gene, age, and gender
P <- list()
# lattice implementation
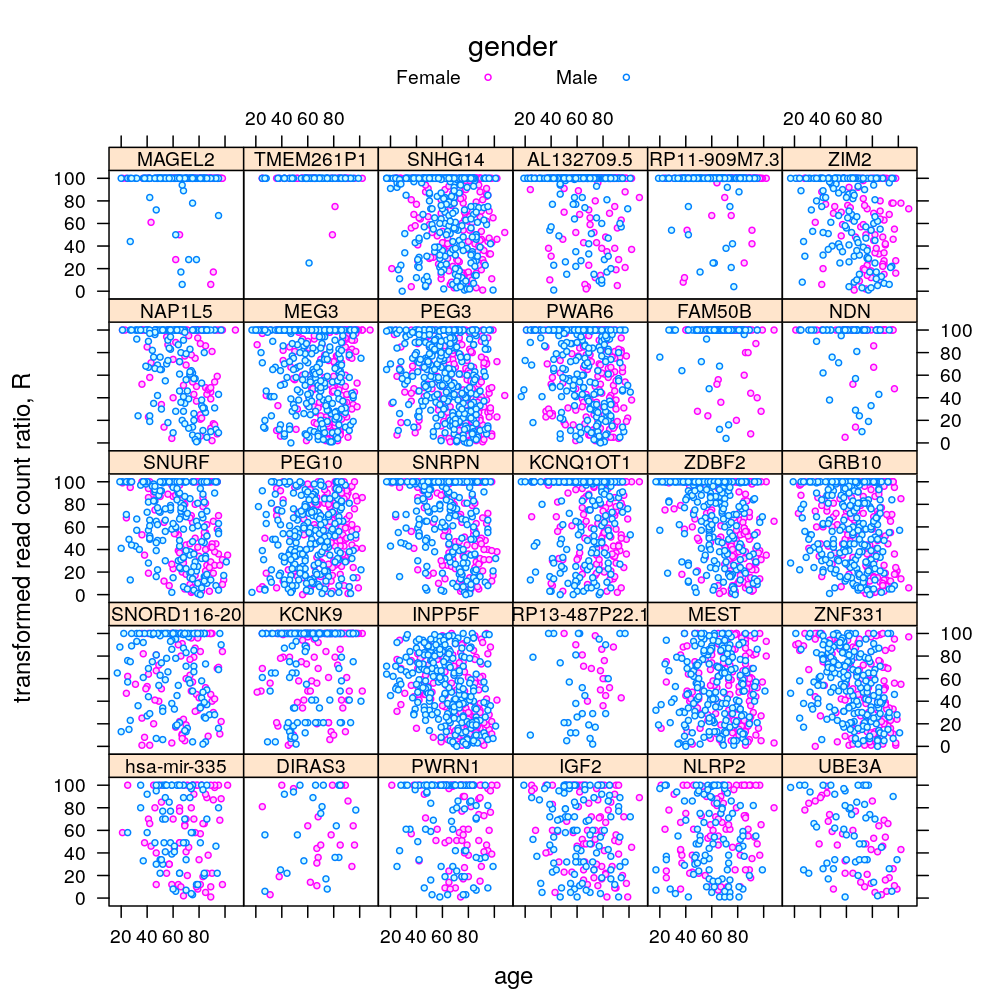
P$s.age.Dx$lattice <-
xyplot(S ~ Age | Gene, data = Y.long, groups = Dx,
subset = Gene %in% gene.ids,
panel = function(x, y, ...) {
panel.xyplot(x, y, pch = 21, ...)
#panel.smoother(x, y, col = "black", lwd = 2, ...)
},
par.settings = list(add.text = list(cex = 0.8),
superpose.symbol = list(cex = 0.5,
fill = trellis.par.get("superpose.symbol")$fill[c(2, 1)],
col = trellis.par.get("superpose.symbol")$col[c(2, 1)])),
auto.key = list(title = "Dx", columns = 2),
ylab = "read count ratio, S",
xlab = "age",
aspect = "fill", layout = c(6, 5))
# ggplot2 implementation
g <- ggplot(data = Y.long, aes(x = Age, y = S))
g <- g + geom_point(pch = "o", aes(color = Dx))
g <- g + geom_smooth(method = "loess", color = "black")
g <- g + facet_wrap(~ Gene)
P$s.age.Dx$ggplot2 <- g
plot(P$s.age.Dx$lattice)
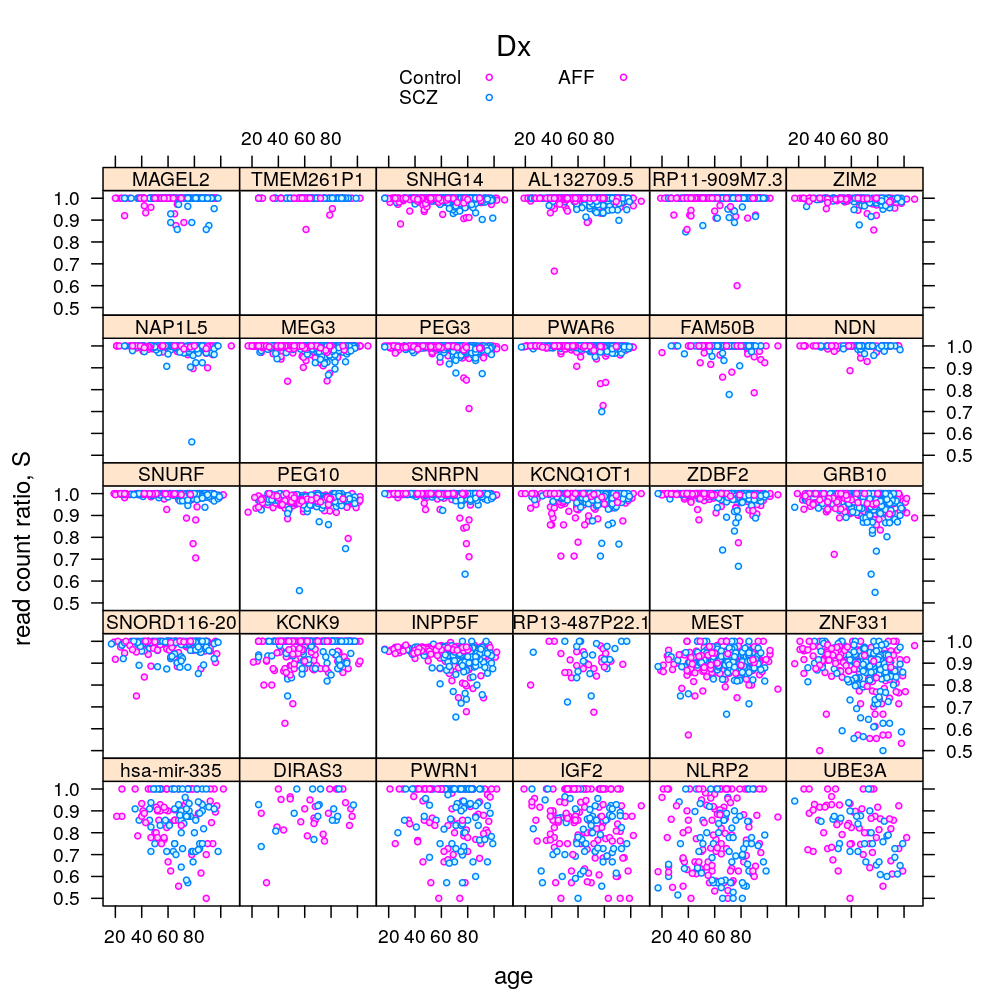
#plot(P$s.age.Dx$ggplot2)
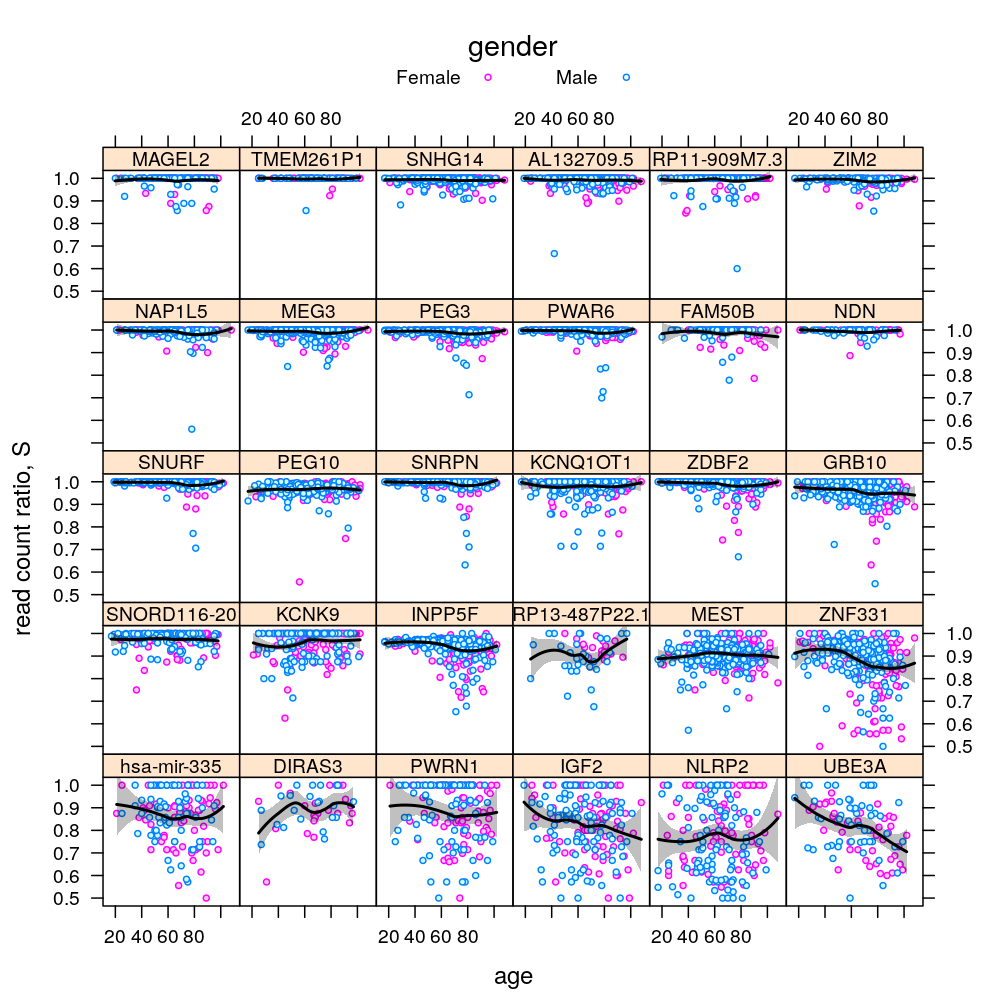
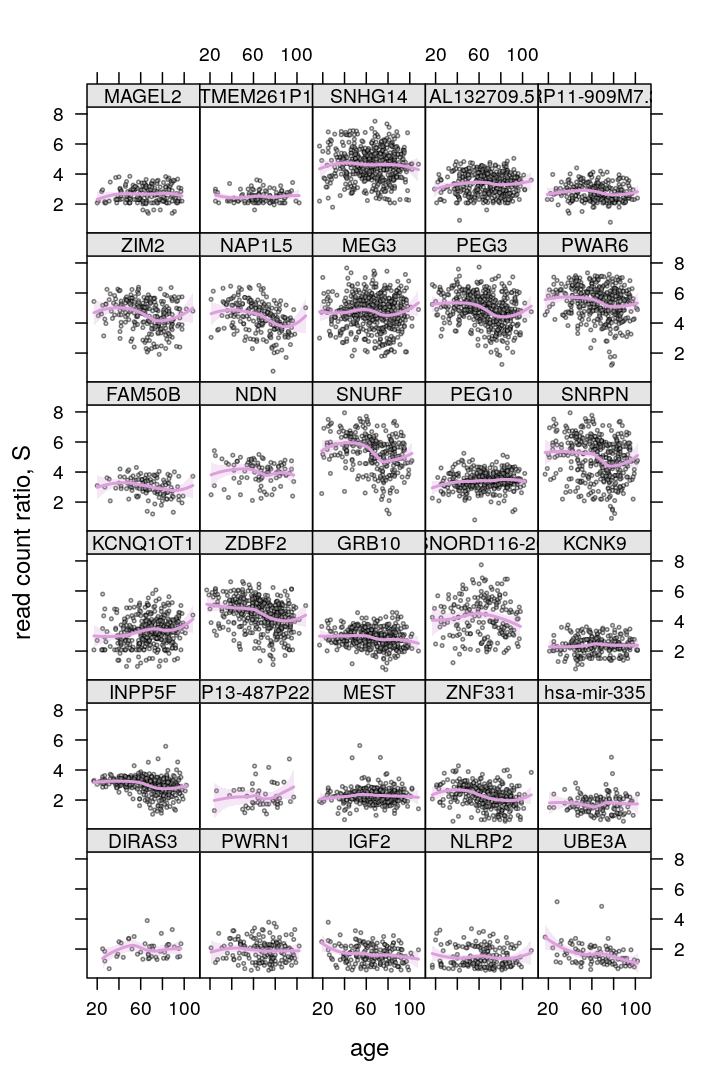
P$s.age$lattice <-
xyplot(S ~ Age | Gene, data = Y.long,
subset = Gene %in% gene.ids,
par.settings = list(add.text = list(cex = 0.8),
strip.background = list(col = "gray90"),
plot.symbol = list(pch = 21, cex = 0.5, col = "black", fill = "gray", alpha = 0.5)),
auto.key = list(title = "gender", columns = 2),
panel = function(x, y, ...) {
panel.xyplot(x, y, pch = 21, cex = 0.3, ...)
panel.smoother(x, y, col = "plum", lwd = 2, ...)
},
ylab = "read count ratio, S",
xlab = "age",
aspect = "fill", layout = c(6, 5))
plot(P$s.age$lattice)
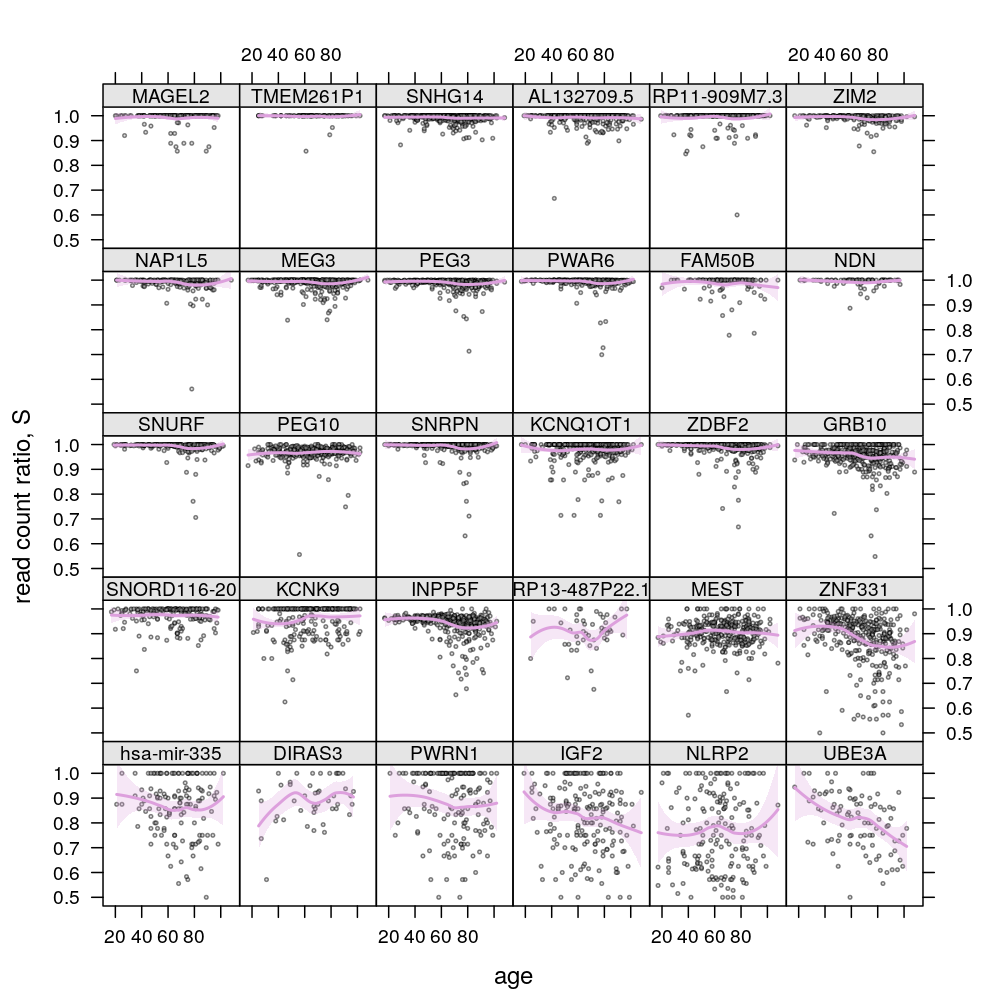
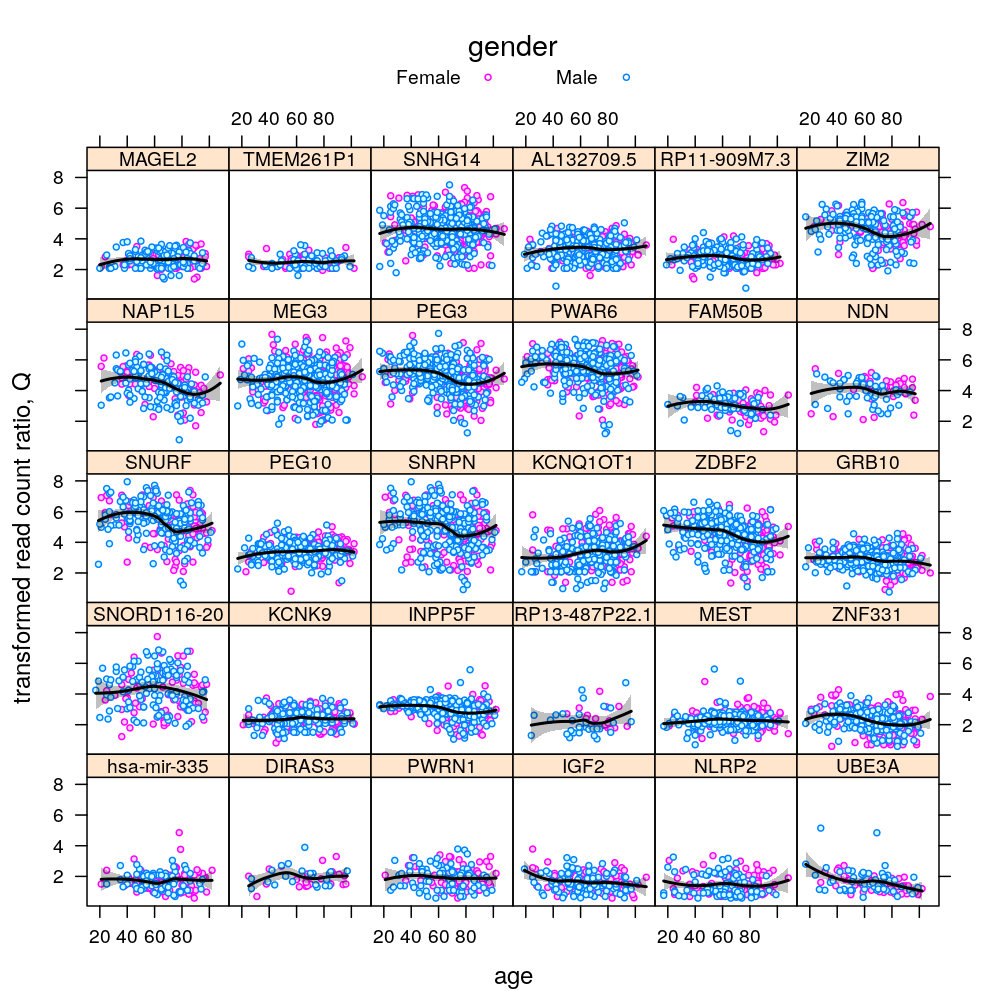
P$s.age$lattice <-
xyplot(Q ~ Age | Gene, data = Y.long,
subset = Gene %in% gene.ids,
par.settings = list(add.text = list(cex = 0.8),
strip.background = list(col = "gray90"),
plot.symbol = list(pch = 21, cex = 0.5, col = "black", fill = "gray", alpha = 0.5)),
auto.key = list(title = "gender", columns = 2),
panel = function(x, y, ...) {
panel.xyplot(x, y, pch = 21, cex = 0.3, ...)
panel.smoother(x, y, col = "plum", lwd = 2, ...)
},
ylab = "read count ratio, S",
xlab = "age",
aspect = "fill", layout = c(6, 5))
plot(P$s.age$lattice)
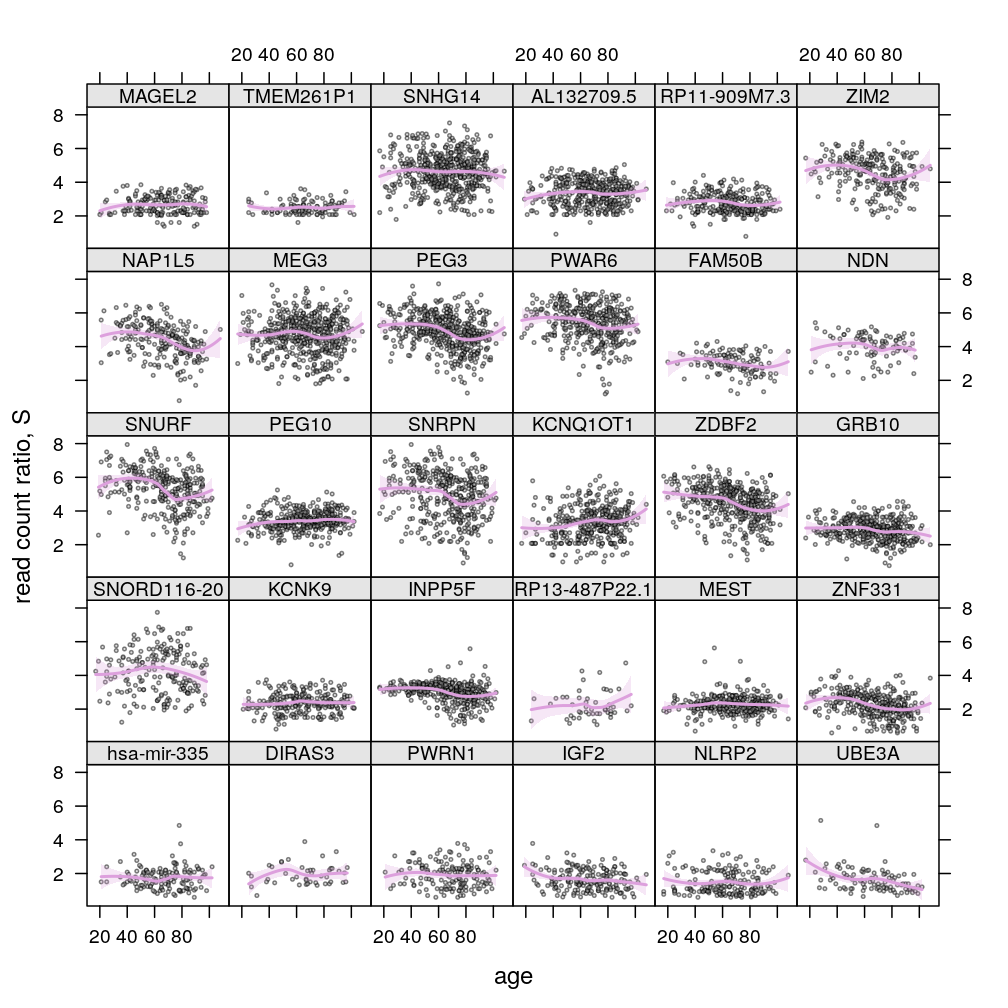
update(P$s.age$lattice, layout = c(5, 6))
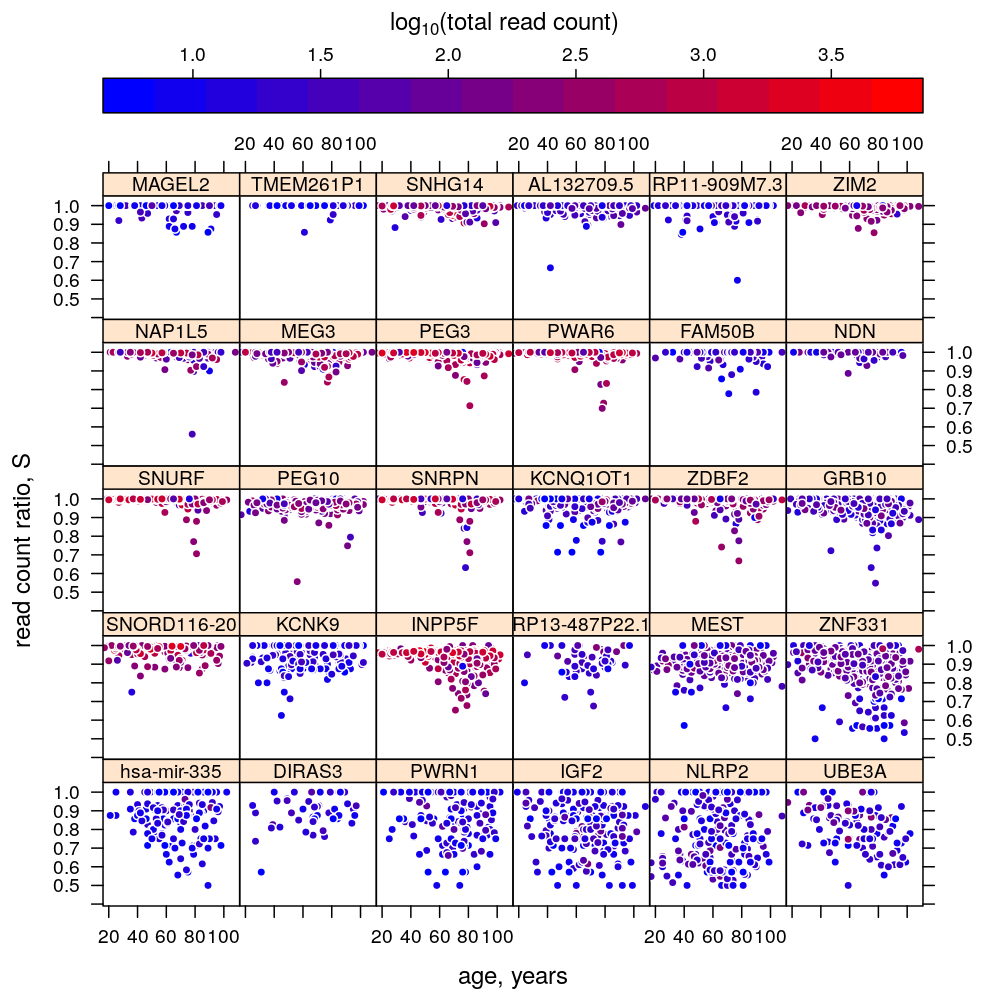
Dependence on gene, age, and total read count

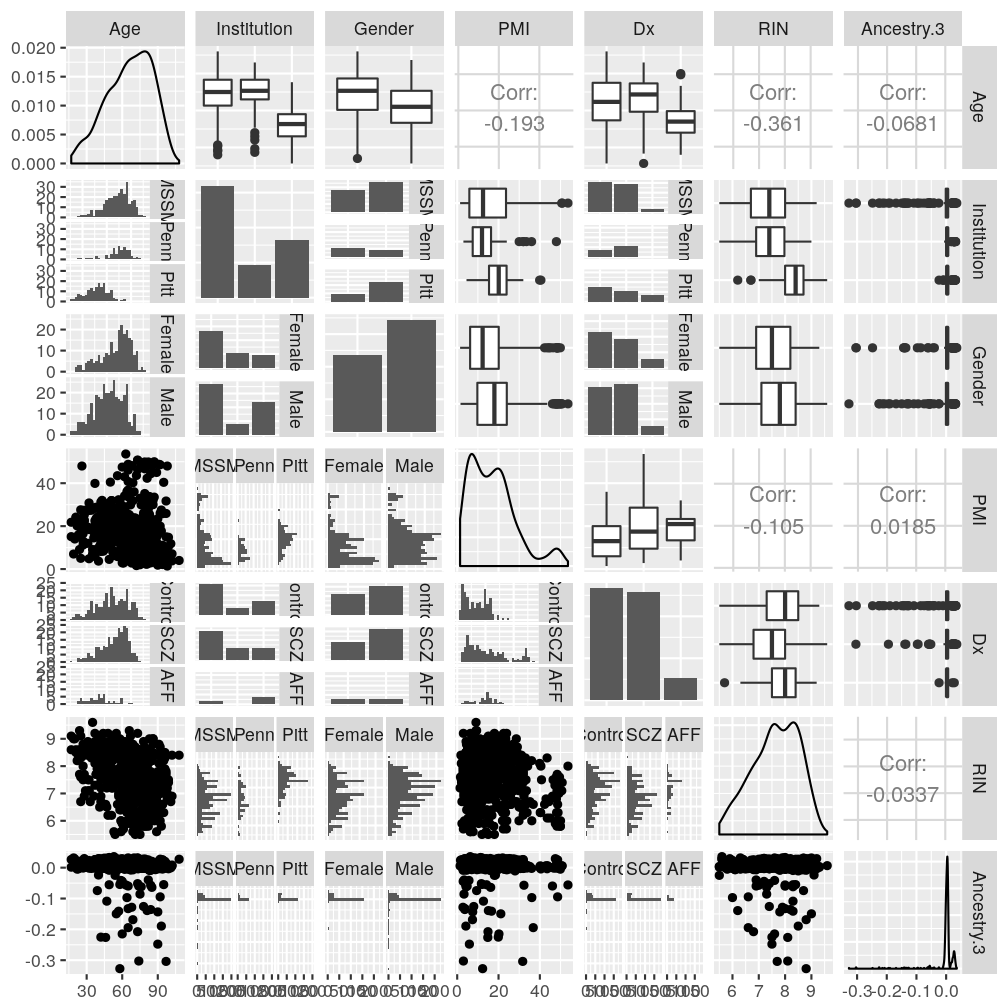
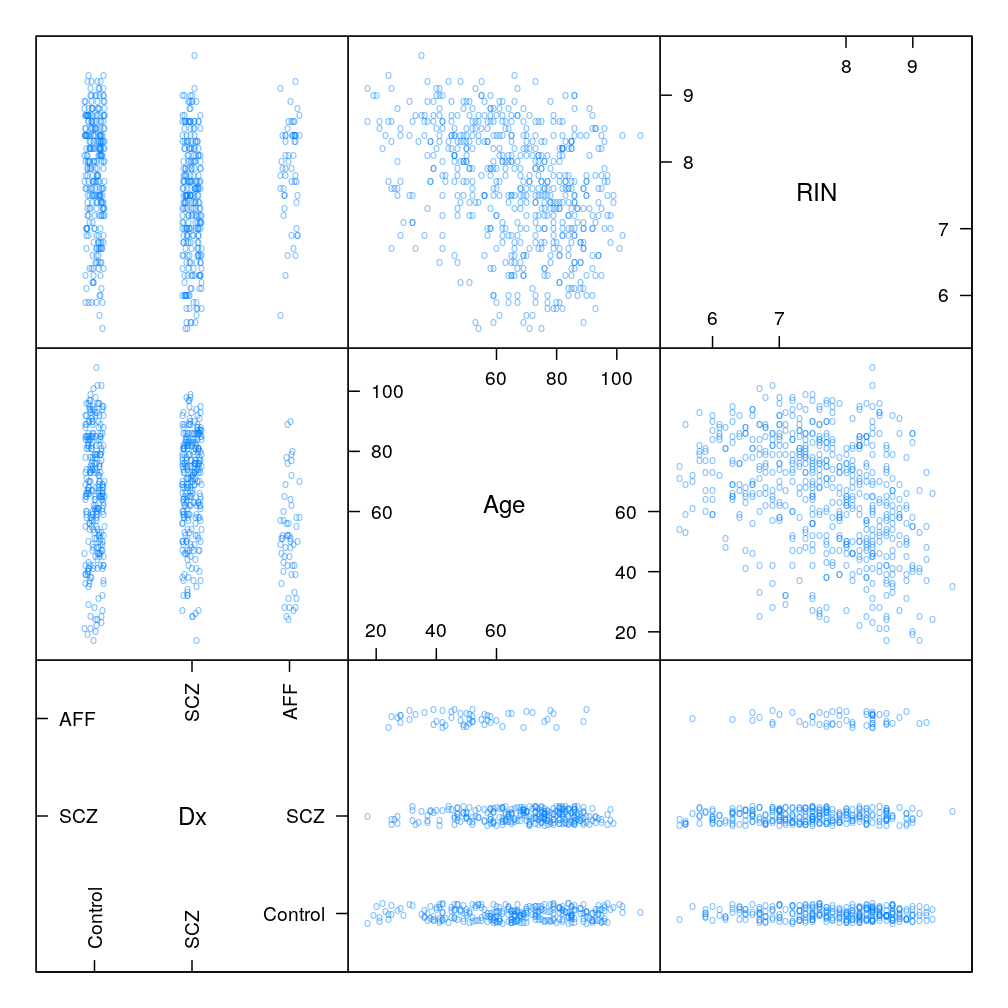
Associations between explanatory variables
Deterministic association: RIN and RIN2
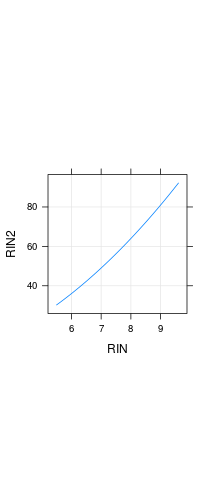

Stochastic (statistical) associations
Both “scatter plot matrices” show the same set of pairwise associations (top: lattice, bottom: ggplot2 and GGally packages).